Gouttenoire and Moradpour Lab

Jérôme Gouttenoire, PhD, PD
Tel. +41 21 314 0749
Mobile: +41 79 556 6093
Contact mail

Darius Moradpour, MD, Professor of Medicine
Tel. +41 21 314 4714
Fax: +41 21 314 4718
Contact mail
Molecular virology and pathogenesis of hepatitis C and E
Research in our laboratory focuses on the molecular basis of liver diseases, especially the virology and pathogenesis of hepatitis C and E. Ongoing projects are funded by the Swiss National Science Foundation and other sources. They involve molecular, biochemical and cell biological as well as virological techniques, including infectious cell culture systems, advanced imaging as well as large-scale gene expression and proteomics analyses. These are complemented on a collaborative basis by genetic and structural analyses.
Hepatitis C
Hepatitis C virus (HCV) infection is a leading cause of chronic hepatitis, liver cirrhosis and hepatocellular carcinoma worldwide. Our research over the past years focused on the viral and cellular determinants required for the formation of a functional HCV replication complex, the investigation of structural and functional properties of the viral proteins, and analyses of interactions of these proteins with cellular proteins and functional pathways.
HCV Interaction with Host Cellular Pathways
HCV has evolved various strategies to counteract the host immune response and to establish persistent infection. Previous work has identified the NS3-4A protease as a key viral protein blocking antiviral innate immune defense as well as other cellular pathways. NS3-4A cleaves and thereby inactivates the essential adaptor molecule MAVS in the RIG-I viral RNA-sensing pathway, thereby blocking interferon production. More recently, we have pursued a quantitative proteomics-based strategy to identify novel cellular targets of the HCV NS3-4A protease. These studies have revealed several novel host targets, including glutathione peroxidase 8 (GPx8), that are currently being characterized for their role in the viral life cycle as well as the pathogenesis of hepatitis C.
Hepatitis E
Hepatitis E virus (HEV) infection is one of the most common causes of acute hepatitis and jaundice in the world. HEV genotypes 1 and 2 cause primarily waterborne outbreaks in low-income countries with poor sanitation while genotypes 3 and 4 have emerged as porcine zoonosis in high-income countries. In addition, HEV genotype 3 can trigger acute-on-chronic liver failure, has been recognized as a cause of diverse neurological complications and can evolve into chronic hepatitis E in immunocompromised patients.
HEV is a positive-strand RNA virus belonging to the Hepeviridae family. Its 7.2-kb RNA genome harbors 3 open reading frames (ORF) that are translated into (i) the ORF1 replicase protein, comprising the RNA-dependent RNA polymerase and other functional domains, (ii) ORF2, the viral capsid, and (iii) ORF3, a small protein required for virion secretion.
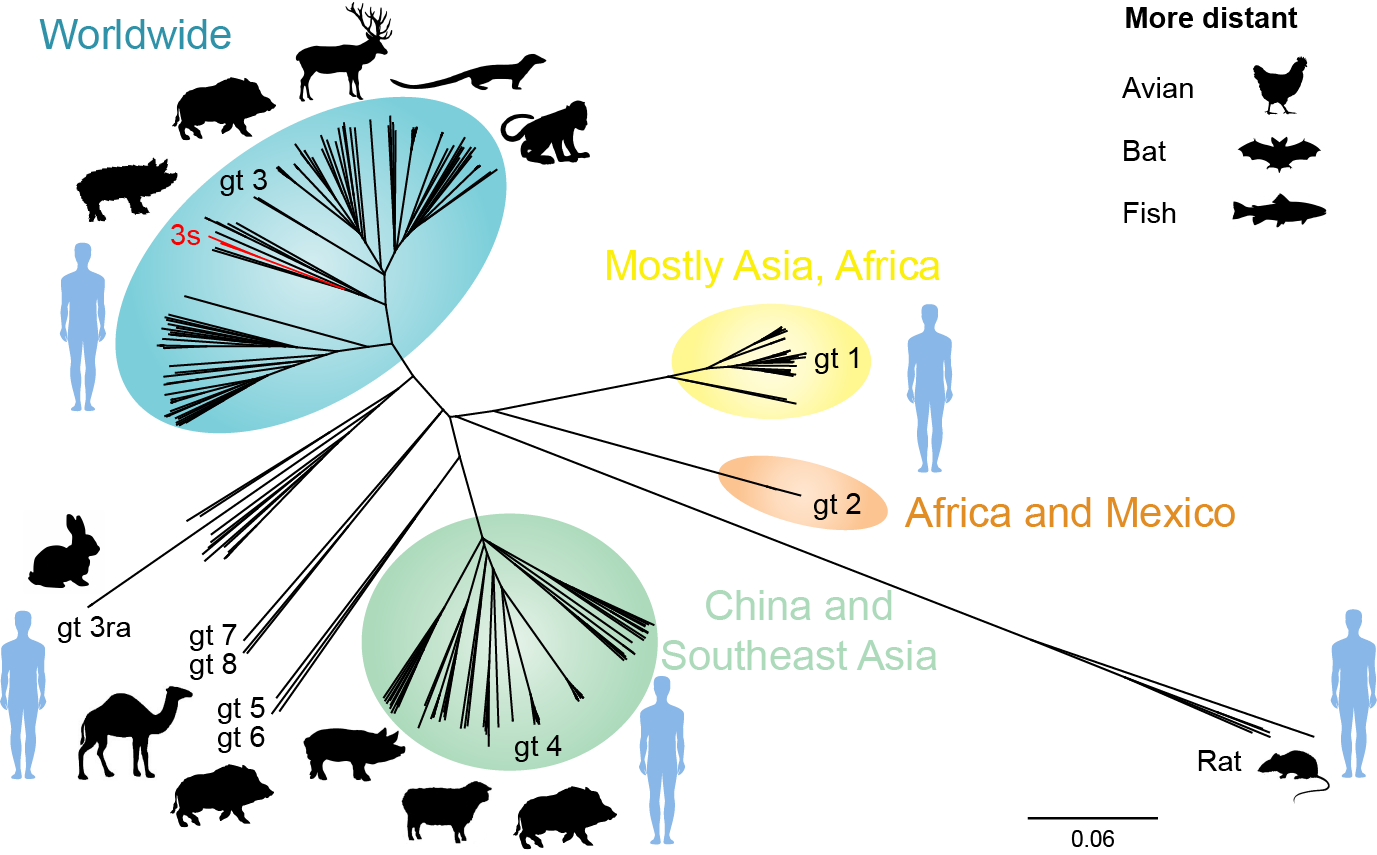
Phylogenetic relationship of hepeviruses identified in different hosts. (updated from Debing Y, Moradpour D, Neyts J and Gouttenoire J. J Hepatol 2016;65:200-212).
Characterization of Hepatitis E in Switzerland
The Service of Gastroenterology and Hepatology in collaboration with different partners at Lausanne University Hospital and the University of Lausanne pursues since a few years a multidisciplinary research effort on hepatitis E, including basic research, diagnostic testing, epidemiology and clinical management. The clinical data and biological samples obtained within the framework of this ongoing work represent an invaluable resource for the studies in our laboratory. In this context, we have recently found that the vast majority of cases of hepatitis E acquired in Switzerland are caused by a particular HEV subtype, genotype 3s, which has been identified only in Switzerland so far. In addition, we have observed that some immunocompromised patients are infected with rabbit HEV. Ongoing projects exploit next generation sequencing and host genetic analyses to correlate viral and host determinants with specific clinical outcomes. Moreover, we are in the process of assembling functional viral clones to further investigate the HEV life cycle and pathogenesis as well as to evaluate antiviral drug candidates.
Exploring the HEV Life Cycle
The HEV life cycle is still poorly characterized. Therefore, our ongoing research on the molecular virology of hepatitis E focuses on the functional organization of the HEV replicase, the structure and function of the ORF3 protein, as well as on the development of novel in vitro model systems.
A transposon-mediated random insertion screen allowed us to identify sites within the HEV ORF1 which tolerate insertions of various tags, such as the HA epitope or a small version of luciferase. These novel tools facilitate our current studies on the subcellular localization and composition of viral replication site and the identification of host factors required for HEV RNA replication. Moreover, the contribution of the host machinery to viral RNA replication is being investigated by the use of CRISPR/Cas9-based genome-wide screening and proteomic approaches.
Our recent work on the HEV ORF3 protein has revealed that palmitoylation is crucial for its stability, membrane association, subcellular localization and essential role in virus production. Further characterization of the ORF3 protein includes structural and biochemical analyses as well as the investigation of the host interacting partners. These ongoing efforts shall further delineate the function(s) of the ORF3 protein in the HEV life cycle.
.



