- Accueil
- Research
- Research Groups
- Gilbert Greub
Gilbert Greub Lab

Gilbert Greub, MD-PhD, FAMH, Full Professor
Tel. +41 21 314 4979
Mobile: +41 79 556 1795
Contact mail
Curriculum Vitae
Full Publications list (01.2021)
UNIL page of Gilbert Greub
Research topics
Discovery of new agents of pneumonia and miscarriage
To isolate new bacterial species, we employ an innovative cell culture system that uses free-living amoebae as cellular background. This amoebal co-culture technique selectively grows amoebae-resisting bacteria, i.e. potential pathogens to which humans are frequently exposed (since amoebae are widespread in water), that are highly resistant to water decontamination, and that resist destruction by human macrophages. Once isolated, the role of each new amoebae-resisting species is tested for its pathogenic potential by comparing seroprevalence in patients with and without pneumonia, and by demonstrating the presence of the bacteria in lower respiratory tract samples taken from patients with pneumonia of unknown etiology. This is achieved by developing new diagnostic approaches for each new species (PCR, improved cell culture, immunohistochemistry). For some selected species, we also develop an animal model of pneumonia. Using a similar approach, we recently identified the role of Waddlia chondrophila in miscarriage.
Study of the biology of Chlamydia-like organisms
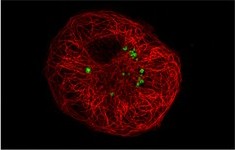
Chlamydia-related bacteria such as Parachlamydia and Waddlia emerge as novel agents of pneumonia and miscarriage. However, their biology is as yet poorly studied. Thus, we started several genomic projects on various Chlamydia-related bacteria, in order to better understand the biology and evolution of these strict intracellular bacteria. We also aim at understanding how these Chlamydia-like bacteria interact with macrophages and at identifying genes that might be involved in the virulence of these obligate intracellular bacteria. Our work mainly focuses on the intracellular trafficking of Chlamydia-like organisms and on their impact on the host microbicidal effectors. Thus, we showed recently that Waddlia is able to recruit mitochondria thank to the actin and microtubule system (Figure).
CHUV
Rue du Bugnon 48
CH-1011 Lausanne, Suisse
+41 21 314 4056



